xarray EOPF backend - Sentinel-3 Native Mode
Learn how to access Sentinel-3 products in native mode

Table of Contents¶
Run this notebook interactively with all dependencies pre-installed
Introduction¶
xarray-eopf is a Python package that extends xarray with a custom backend called "eopf-zarr". This backend enables seamless access to ESA EOPF data products stored in the Zarr format, presenting them as analysis-ready data structures.
In this notebook, we demonstrate how to use the xarray-eopf backend to access Sentinel-3 EOPF Zarr products in native mode. All data access is lazy, meaning that data is only loaded when required—for example, during plotting or when writing to storage.
For a general introduction to the xarray EOPF backend, see the introduction notebook.
🐙 GitHub: EOPF Sample Service – xarray-eopf
❗ Issue Tracker: Submit or view issues
📘 Documentation: xarray-eopf Docs
Import Modules¶
The xarray-eopf backend is implemented as a plugin for xarray. Once installed, it registers automatically and requires no additional import. You can simply import xarray as usual:
import datetime
import matplotlib.pyplot as plt
import pystac_client
import xarray as xrOpen a Sentinel-3 OLCI L1 EFR¶
We begin with an example that accesses a Sentinel-3 OLCI L1 EFR product in native mode.
Find a Sentinel-3 OLCI L1 EFR Zarr Sample via STAC¶
To obtain a product URL, you can use the STAC Browser to search for a Sentinel-3 OLCI Level-1 EFR tile.
catalog = pystac_client.Client.open("https://stac.core.eopf.eodc.eu")
items = list(
catalog.search(
collections=["sentinel-3-olci-l1-efr"],
bbox=[7.2, 44.5, 7.4, 44.7],
datetime=[str(datetime.date.today() - datetime.timedelta(days=5)), None],
).items()
)
items[<Item id=S3B_OL_1_EFR____20260329T094553_20260329T094853_20260329T114455_0179_118_193_2160_ESA_O_NR_004>,
<Item id=S3B_OL_1_EFR____20260328T101204_20260328T101504_20260328T121637_0179_118_179_2160_ESA_O_NR_004>,
<Item id=S3B_OL_1_EFR____20260328T101204_20260328T101504_20260329T100706_0180_118_179_2160_ESA_O_NT_004>,
<Item id=S3A_OL_1_EFR____20260327T093557_20260327T093857_20260328T103721_0179_137_307_2160_PS1_O_NT_004>,
<Item id=S3A_OL_1_EFR____20260327T093557_20260327T093857_20260327T113653_0180_137_307_2160_PS1_O_NR_004>,
<Item id=S3A_OL_1_EFR____20260326T100207_20260326T100507_20260327T110434_0180_137_293_2160_PS1_O_NT_004>,
<Item id=S3A_OL_1_EFR____20260326T100207_20260326T100507_20260326T120302_0179_137_293_2160_PS1_O_NR_004>,
<Item id=S3B_OL_1_EFR____20260326T092326_20260326T092626_20260327T093749_0179_118_150_2160_ESA_O_NT_004>,
<Item id=S3B_OL_1_EFR____20260326T092326_20260326T092626_20260326T113241_0179_118_150_2160_ESA_O_NR_004>,
<Item id=S3B_OL_1_EFR____20260325T094937_20260325T095237_20260326T102555_0179_118_136_2160_ESA_O_NT_004>,
<Item id=S3B_OL_1_EFR____20260325T094937_20260325T095237_20260325T125634_0179_118_136_2160_ESA_O_NR_004>]Next, we can inspect the item’s contents. The asset "product" links to the entire Zarr store. The additional field xarray:open_datatree_kwargs has been included in the asset "product", which provides the arguments needed to open the product using Xarray’s eopf-zarr engine.
item = items[1]
itemOpen Sentinel-3 OLCI L1 EFR as DataTree¶
We can use the "product" asset to obtain the href and xarray:open_datatree_kwargs from the STAC item, and open the product as an xarray.DataTree as shown below:
dt = xr.open_datatree(
item.assets["product"].href,
**item.assets["product"].extra_fields["xarray:open_datatree_kwargs"],
)
dtAs an example, we plot the red band (oa08_radiance), which will trigger loading and visualization of the data. We also plot the 2D curvilinear latitude and longitude grids, which can be used for geolocation.
Note: To speed up rendering of large datasets in Matplotlib, we plot the data at a lower resolution (every 4th pixel).
fig, ax = plt.subplots(1, 3, figsize=(15, 4))
dt.measurements.oa08_radiance[::4, ::4].plot.imshow(ax=ax[0], vmin=0, vmax=300)
dt.measurements.latitude[::4, ::4].plot.imshow(ax=ax[1], cmap="viridis")
dt.measurements.longitude[::4, ::4].plot.imshow(ax=ax[2], cmap="viridis")
plt.tight_layout()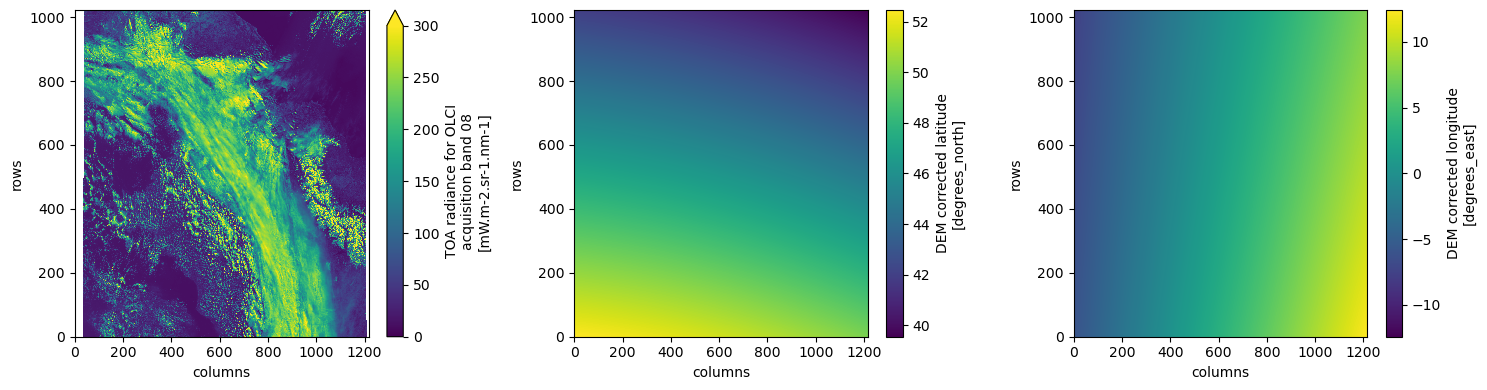
Open Sentinel-3 OLCI L1 EFR Radiance Group as Dataset¶
We can directly access the individual reflectance group (radianceData) as xarray.Dataset objects. The opening parameters are stored in the asset’s extra field "xarray:open_dataset_kwargs".
ds = xr.open_dataset(
item.assets["radianceData"].href,
**item.assets["radianceData"].extra_fields["xarray:open_dataset_kwargs"],
)
dsWe can also filter the varaibles by band names as shown below:
ds = xr.open_dataset(
item.assets["radianceData"].href,
**item.assets["radianceData"].extra_fields["xarray:open_dataset_kwargs"],
variables="oa0[468]",
)
dsAnd we can again plot the red band, which triggers loading the data.
ds.oa08_radiance[::4, ::4].plot(vmax=300)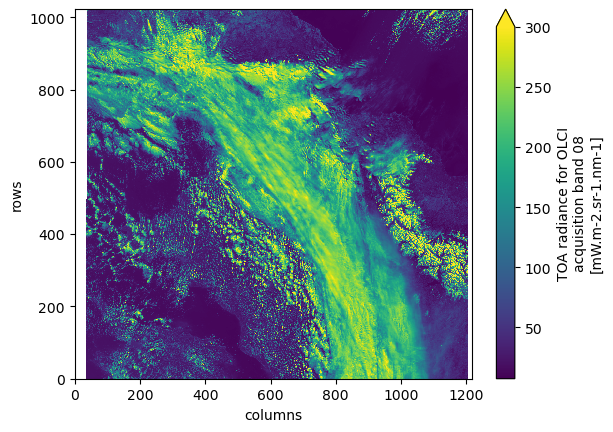
Open Sentinel-3 OLCI L1 EFR as Dataset¶
The xarray.DataTree model was introduced in xarray v2024.10.0 (October 2024). To maintain compatibility with workflows based on xr.Dataset, the function xarray.open_dataset(..., engine="eopf-zarr", op_mode="native") is provided, which flattens the DataTree into a single dataset.
During this process, hierarchical groups in the Zarr product are merged, and variable as well as dimension names are prefixed with their group paths (using _ by default) to ensure uniqueness. For example, measurements/oa08_radiance becomes measurements_oa08_radiance.
ds = xr.open_dataset(
item.assets["product"].href,
engine="eopf-zarr",
op_mode="native",
chunks={},
)
dsThe separator character used in flattened variable names can be customized via the group_sep parameter. Additionally, you can filter the returned variables using the variables keyword argument, which accepts a string, an iterable of names, or a regular expression (regex) pattern.
ds = xr.open_dataset(
item.assets["product"].href,
engine="eopf-zarr",
op_mode="native",
chunks={},
group_sep="/",
variables="measurements*.",
)
dsAlso here, we can plot one spectral band as an example.
ds["measurements/oa08_radiance"][::4, ::4].plot(vmin=0.0, vmax=300.0)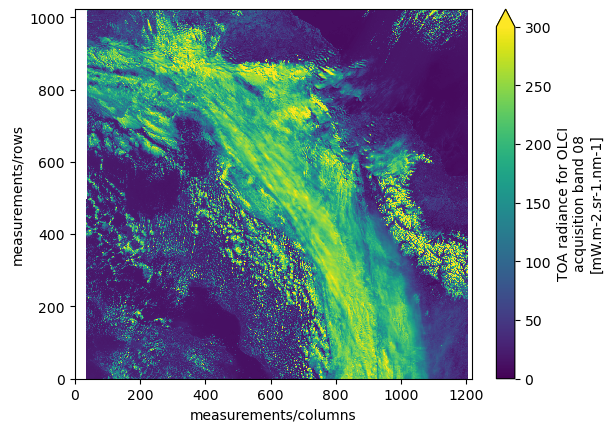
Open a Sentinel-3 OLCI L1 ERR¶
We now access a Sentinel-3 OLCI L1 ERR product in native mode. The data access methods shown above apply equally to Sentinel-3 OLCI L1 EFR.
Find a Sentinel-3 OLCI L1 ERR Zarr Sample via STAC¶
To obtain a product URL, you can use the STAC Browser to search for available Sentinel-3 OLCI L1 ERR tiles.
catalog = pystac_client.Client.open("https://stac.core.eopf.eodc.eu")
items = list(
catalog.search(
collections=["sentinel-3-olci-l1-err"],
bbox=[7.2, 44.5, 7.4, 44.7],
datetime=[str(datetime.date.today() - datetime.timedelta(days=5)), None],
).items()
)
items[<Item id=S3B_OL_1_ERR____20260329T093543_20260329T101948_20260329T120344_2645_118_193______ESA_O_NR_004>,
<Item id=S3B_OL_1_ERR____20260328T100201_20260328T104605_20260329T095351_2644_118_179______ESA_O_NT_004>,
<Item id=S3B_OL_1_ERR____20260328T100201_20260328T104605_20260328T122605_2644_118_179______ESA_O_NR_004>,
<Item id=S3A_OL_1_ERR____20260327T092601_20260327T101004_20260328T103618_2643_137_307______PS1_O_NT_004>,
<Item id=S3A_OL_1_ERR____20260327T092601_20260327T101004_20260327T113524_2643_137_307______PS1_O_NR_004>,
<Item id=S3A_OL_1_ERR____20260326T095219_20260326T103622_20260327T110333_2643_137_293______PS1_O_NT_004>,
<Item id=S3A_OL_1_ERR____20260326T095219_20260326T103622_20260326T120200_2643_137_293______PS1_O_NR_004>,
<Item id=S3B_OL_1_ERR____20260326T091338_20260326T095741_20260327T092230_2643_118_150______ESA_O_NT_004>,
<Item id=S3B_OL_1_ERR____20260326T091338_20260326T095741_20260326T114137_2643_118_150______ESA_O_NR_004>,
<Item id=S3B_OL_1_ERR____20260325T093956_20260325T102359_20260326T101436_2643_118_136______ESA_O_NT_004>,
<Item id=S3B_OL_1_ERR____20260325T093956_20260325T102359_20260325T120957_2643_118_136______ESA_O_NR_004>]item = items[0]
itemOpen Sentinel-3 OLCI L1 ERR as DataTree¶
We can use the "product" asset to obtain the href and xarray:open_datatree_kwargs from the STAC item, and open the product as an xarray.DataTree as shown below:
dt = xr.open_datatree(
item.assets["product"].href,
**item.assets["product"].extra_fields["xarray:open_datatree_kwargs"],
)
dtOpen Sentinel-3 OLCI L1 EFF Radiance Group as Dataset¶
We can directly access the individual reflectance group (radianceData) as xarray.Dataset objects. The opening parameters are stored in the asset’s extra field "xarray:open_dataset_kwargs".
ds = xr.open_dataset(
item.assets["radianceData"].href,
**item.assets["radianceData"].extra_fields["xarray:open_dataset_kwargs"],
)
dsAs an example, we plot the red band (oa08_radiance), which will trigger loading and visualization of the data. We also plot the 2D curvilinear latitude and longitude grids, which can be used for geolocation. Note that one product spanns half an orbit.
Note: To speed up rendering of large datasets in Matplotlib, we plot the data at a lower resolution (every 4th pixel along-track).
fig, ax = plt.subplots(1, 3, figsize=(10, 5))
ds.oa08_radiance[::4, :].plot.imshow(ax=ax[0], vmin=0, vmax=300)
ds.latitude[::4, :].plot.imshow(ax=ax[1], cmap="viridis")
ds.longitude[::4, :].plot.imshow(ax=ax[2], cmap="viridis")
plt.tight_layout()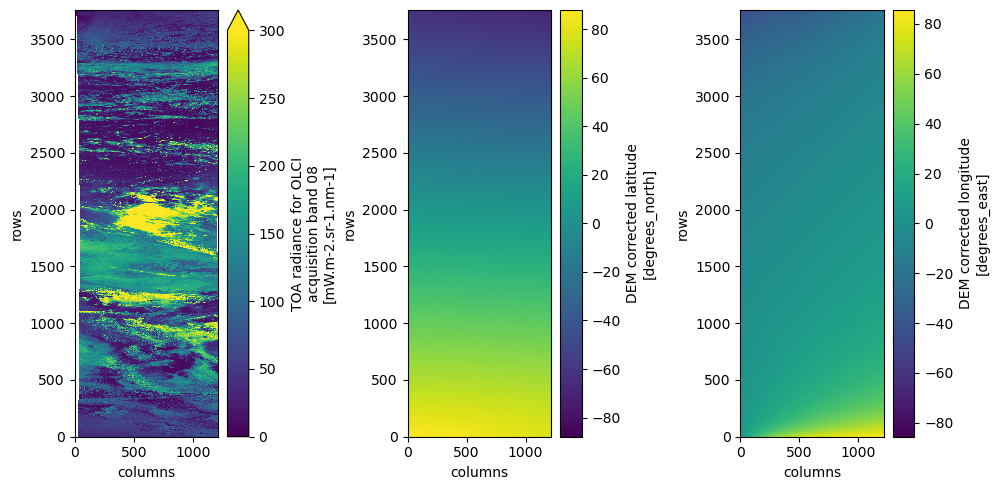
Open a Sentinel-3 SLSTR Level-2 LST¶
We now access a Sentinel-3 SLSTR Level-2 LST product in native mode. The data access methods shown above apply equally to Sentinel-3 SLSTR Level-2 LST.
Find a Sentinel-3 SLSTR Level-2 LST Zarr Sample via STAC¶
To obtain a product URL, you can use the STAC Browser to search for a Sentinel-3 SLSTR Level-2 LST tile.
catalog = pystac_client.Client.open("https://stac.core.eopf.eodc.eu")
items = list(
catalog.search(
collections=["sentinel-3-slstr-l2-lst"],
bbox=[7.2, 44.5, 7.4, 44.7],
datetime=[str(datetime.date.today() - datetime.timedelta(days=5)), None],
).items()
)
items[<Item id=S3A_SL_2_LST____20260329T214729_20260329T215029_20260330T003114_0179_137_343_0720_PS1_O_NR_004>,
<Item id=S3B_SL_2_LST____20260329T210848_20260329T211148_20260329T235533_0179_118_200_0720_ESA_O_NR_004>,
<Item id=S3A_SL_2_LST____20260329T102435_20260329T102735_20260329T125030_0179_137_336_2160_PS1_O_NR_004>,
<Item id=S3B_SL_2_LST____20260329T094553_20260329T094853_20260329T121220_0179_118_193_2160_ESA_O_NR_004>,
<Item id=S3B_SL_2_LST____20260328T213459_20260328T213759_20260329T001031_0179_118_186_0720_ESA_O_NR_004>,
<Item id=S3B_SL_2_LST____20260328T213459_20260328T213759_20260330T051647_0179_118_186_0720_ESA_O_NT_004>,
<Item id=S3A_SL_2_LST____20260328T203241_20260328T203541_20260328T231506_0179_137_328_0720_PS1_O_NR_004>,
<Item id=S3A_SL_2_LST____20260328T203241_20260328T203541_20260330T081257_0179_137_328_0720_PS1_O_NT_004>,
<Item id=S3B_SL_2_LST____20260328T101204_20260328T101504_20260328T123424_0179_118_179_2160_ESA_O_NR_004>,
<Item id=S3B_SL_2_LST____20260328T101204_20260328T101504_20260329T172450_0180_118_179_2160_ESA_O_NT_004>,
<Item id=S3A_SL_2_LST____20260327T205851_20260327T210151_20260329T083311_0179_137_314_0720_PS1_O_NT_004>,
<Item id=S3A_SL_2_LST____20260327T205851_20260327T210151_20260327T234107_0179_137_314_0720_PS1_O_NR_004>,
<Item id=S3A_SL_2_LST____20260327T093557_20260327T093857_20260328T204053_0179_137_307_2160_PS1_O_NT_004>,
<Item id=S3A_SL_2_LST____20260327T093557_20260327T093857_20260327T120121_0180_137_307_2160_PS1_O_NR_004>,
<Item id=S3A_SL_2_LST____20260326T212502_20260326T212802_20260328T091525_0179_137_300_0720_PS1_O_NT_004>,
<Item id=S3A_SL_2_LST____20260326T212502_20260326T212802_20260327T000636_0179_137_300_0720_PS1_O_NR_004>,
<Item id=S3B_SL_2_LST____20260326T204621_20260326T204921_20260328T030457_0179_118_157_0720_ESA_O_NT_004>,
<Item id=S3B_SL_2_LST____20260326T204621_20260326T204921_20260326T232930_0180_118_157_0720_ESA_O_NR_004>,
<Item id=S3A_SL_2_LST____20260326T100207_20260326T100507_20260327T212028_0180_137_293_2160_PS1_O_NT_004>,
<Item id=S3A_SL_2_LST____20260326T100207_20260326T100507_20260326T122451_0179_137_293_2160_PS1_O_NR_004>,
<Item id=S3B_SL_2_LST____20260326T092326_20260326T092626_20260327T153634_0179_118_150_2160_ESA_O_NT_004>,
<Item id=S3B_SL_2_LST____20260326T092326_20260326T092626_20260326T114832_0179_118_150_2160_ESA_O_NR_004>,
<Item id=S3B_SL_2_LST____20260325T211231_20260325T211531_20260327T033756_0179_118_143_0720_ESA_O_NT_004>,
<Item id=S3B_SL_2_LST____20260325T211231_20260325T211531_20260325T234726_0179_118_143_0720_ESA_O_NR_004>,
<Item id=S3A_SL_2_LST____20260325T102818_20260325T103118_20260326T221710_0180_137_279_2160_PS1_O_NT_004>,
<Item id=S3A_SL_2_LST____20260325T102818_20260325T103118_20260325T125251_0180_137_279_2160_PS1_O_NR_004>,
<Item id=S3B_SL_2_LST____20260325T094937_20260325T095237_20260326T161103_0179_118_136_2160_ESA_O_NT_004>,
<Item id=S3B_SL_2_LST____20260325T094937_20260325T095237_20260325T122544_0179_118_136_2160_ESA_O_NR_004>]item = items[0]
itemOpen Sentinel-3 SLSTR Level-2 LST as DataTree¶
We can use the "product" asset to obtain the href and xarray:open_datatree_kwargs from the STAC item, and open the product as an xarray.DataTree as shown below:
dt = xr.open_datatree(
item.assets["product"].href,
**item.assets["product"].extra_fields["xarray:open_datatree_kwargs"],
)
dtOpen Sentinel-3 SLSTR Level-2 LST Group as Dataset¶
Similarly, we can directly access the individual LST group (lst) as xarray.Dataset objects.
ds = xr.open_dataset(
item.assets["lst"].href, engine="eopf-zarr", op_mode="native", chunks={}
)
dsAnd we can plot the LST array along with the 2D latitude and longitude grids.
fig, ax = plt.subplots(1, 3, figsize=(15, 4))
ds.lst.plot.imshow(ax=ax[0])
ds.latitude.plot.imshow(ax=ax[1], cmap="viridis")
ds.longitude.plot.imshow(ax=ax[2], cmap="viridis")
plt.tight_layout()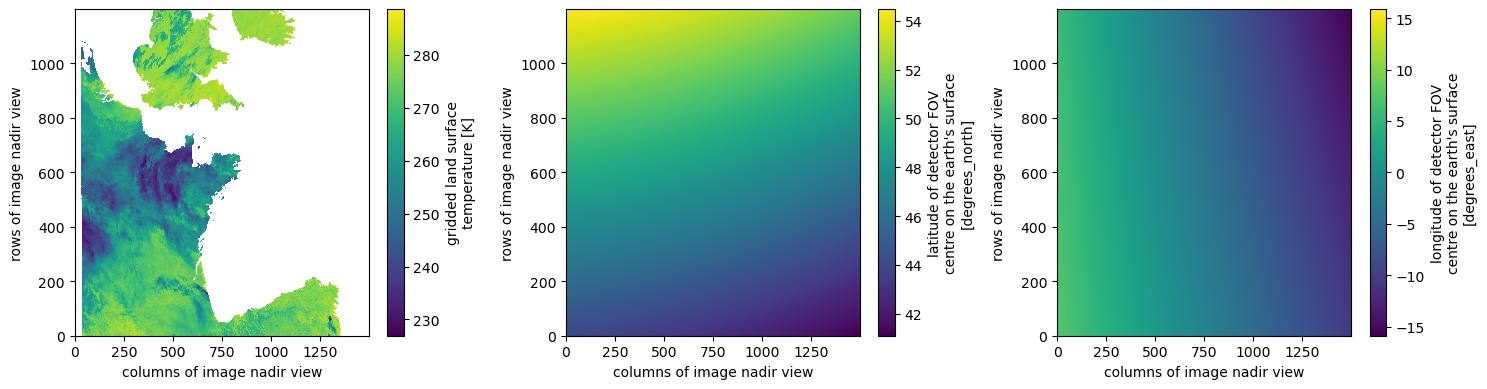
Other Sentinel-3 products can be accessed using the same approach and are therefore not shown here in detail.
Conclusion¶
This notebook demonstrates how to access Sentinel-3 EOPF Zarr samples in native mode using the xarray-eopf plugin. Key takeaways are:
Access all Sentinel-3 products using hte same methods.
Open the full Zarr store as an
xr.DataTreeusingxr.open_datasetand the asset"product".Open subgroups (e.g.,
radianceData,lst) asxr.Datasetusingxr.open_dataset.Open the full Zarr store as a flattened
xr.Datasetusingxr.open_datasetand the asset"product".Filter variables using the
variableskeyword argument.Opening parameters are integrated in STAC assets.
Note: This notebook only covers the native mode, which presents the data as close as possible to the original product.
For an analysis-ready view, see the Sentinel-3 analysis mode notebook.
