Sentinel-3 Patchwork L1 OLCI EFR L2 SLSTR FPR SYN AOD product format prototype
Sentinel-3 L1 OLCI EFR, Sentinel-3 L2 SLSTR FPR, Sentinel-3 SYN AOD product
Table of Contents¶
Run this notebook interactively with all dependencies pre-installed
Introduction¶
In this notebook we will show an example of using Sentinel-3 L1 OLCI EFR, Sentinel-3 L2 SLSTR FPR, Sentinel-3 SYN AOD product
import matplotlib.pyplot as plt
import cartopy.crs as ccrs
import fsspec # For remote access
import datatreeL1 OLCI EFR¶
Product can only be accessed locally and also remotely, even if it is a zarr.zip
# Define the remote product path
remote_product_path = "https://eopf-public.s3.sbg.perf.cloud.ovh.net/eoproducts/S03OLCEFR_20230506T015316_0180_B117_T883.zarr.zip"
# Construct the fsspec mapper path
store = fsspec.get_mapper(f"zip::{remote_product_path}")
# Load it with datatree
dt = datatree.open_datatree(store, engine="zarr", consolidated=False, chunks={})Open the product using EOPF API and visualize the tree structure¶
print(dt)DataTree('None', parent=None)
│ Dimensions: ()
│ Data variables:
│ *empty*
│ Attributes:
│ other_metadata: {'absolute_pass_number': 52368, 'cycle_number': 79, 'dat...
│ stac_discovery: {'assets': [], 'bbox': [132.954, 29.0715, 115.986, 41.97...
├── DataTree('conditions')
│ ├── DataTree('geometry')
│ │ Dimensions: (tp_rows: 4092, tp_columns: 77)
│ │ Coordinates:
│ │ latitude (tp_rows, tp_columns) float64 3MB dask.array<chunksize=(4092, 77), meta=np.ndarray>
│ │ longitude (tp_rows, tp_columns) float64 3MB dask.array<chunksize=(4092, 77), meta=np.ndarray>
│ │ Dimensions without coordinates: tp_rows, tp_columns
│ │ Data variables:
│ │ oaa (tp_rows, tp_columns) float64 3MB dask.array<chunksize=(4092, 77), meta=np.ndarray>
│ │ oza (tp_rows, tp_columns) float64 3MB dask.array<chunksize=(4092, 77), meta=np.ndarray>
│ │ saa (tp_rows, tp_columns) float64 3MB dask.array<chunksize=(4092, 77), meta=np.ndarray>
│ │ sza (tp_rows, tp_columns) float64 3MB dask.array<chunksize=(4092, 77), meta=np.ndarray>
│ ├── DataTree('image')
│ │ Dimensions: (rows: 4092, columns: 4865)
│ │ Coordinates:
│ │ altitude (rows, columns) float32 80MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ │ latitude (rows, columns) float64 159MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ │ longitude (rows, columns) float64 159MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ │ Dimensions without coordinates: rows, columns
│ │ Data variables:
│ │ detector_index (rows, columns) float32 80MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ │ frame_offset (rows, columns) float32 80MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ ├── DataTree('instrument')
│ │ Dimensions: (bands: 21, detectors: 3700, bands1: 21,
│ │ bands2: 21)
│ │ Dimensions without coordinates: bands, detectors, bands1, bands2
│ │ Data variables:
│ │ fwhm (bands, detectors) float32 311kB dask.array<chunksize=(21, 3700), meta=np.ndarray>
│ │ lambda0 (bands, detectors) float32 311kB dask.array<chunksize=(21, 3700), meta=np.ndarray>
│ │ relative_spectral_covariance (bands1, bands2) float32 2kB dask.array<chunksize=(21, 21), meta=np.ndarray>
│ │ solar_flux (bands, detectors) float32 311kB dask.array<chunksize=(21, 3700), meta=np.ndarray>
│ ├── DataTree('meteorology')
│ │ Dimensions: (tp_rows: 4092, tp_columns: 77,
│ │ pressure_level: 25, wind_vector: 2)
│ │ Coordinates:
│ │ * pressure_level (pressure_level) float32 100B 1e+03 ... 1.0
│ │ Dimensions without coordinates: tp_rows, tp_columns, wind_vector
│ │ Data variables:
│ │ atmospheric_temperature_profile (tp_rows, tp_columns, pressure_level) float32 32MB dask.array<chunksize=(4092, 77, 25), meta=np.ndarray>
│ │ horizontal_wind (tp_rows, tp_columns, wind_vector) float32 3MB dask.array<chunksize=(4092, 77, 2), meta=np.ndarray>
│ │ humidity (tp_rows, tp_columns) float32 1MB dask.array<chunksize=(4092, 77), meta=np.ndarray>
│ │ sea_level_pressure (tp_rows, tp_columns) float32 1MB dask.array<chunksize=(4092, 77), meta=np.ndarray>
│ │ total_columnar_water_vapour (tp_rows, tp_columns) float32 1MB dask.array<chunksize=(4092, 77), meta=np.ndarray>
│ │ total_ozone (tp_rows, tp_columns) float32 1MB dask.array<chunksize=(4092, 77), meta=np.ndarray>
│ └── DataTree('orphans')
│ Dimensions: (rows: 4092, removed_pixels: 150)
│ Coordinates:
│ altitude (rows, removed_pixels) float32 2MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
│ latitude (rows, removed_pixels) float64 5MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
│ longitude (rows, removed_pixels) float64 5MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
│ Dimensions without coordinates: rows, removed_pixels
│ Data variables:
│ detector_index (rows, removed_pixels) float32 2MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
│ frame_offset (rows, removed_pixels) float32 2MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
│ nb_removed_pixels (rows) uint16 8kB dask.array<chunksize=(1024,), meta=np.ndarray>
│ sza (rows, removed_pixels) float64 5MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
├── DataTree('measurements')
│ │ Dimensions: (rows: 4092, columns: 4865)
│ │ Coordinates:
│ │ latitude (rows, columns) float64 159MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ │ longitude (rows, columns) float64 159MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ │ time_stamp (rows) datetime64[ns] 33kB dask.array<chunksize=(1024,), meta=np.ndarray>
│ │ Dimensions without coordinates: rows, columns
│ │ Data variables: (12/21)
│ │ oa01_radiance (rows, columns) float64 159MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ │ oa02_radiance (rows, columns) float64 159MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ │ oa03_radiance (rows, columns) float64 159MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ │ oa04_radiance (rows, columns) float64 159MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ │ oa05_radiance (rows, columns) float64 159MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ │ oa06_radiance (rows, columns) float64 159MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ │ ... ...
│ │ oa16_radiance (rows, columns) float64 159MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ │ oa17_radiance (rows, columns) float64 159MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ │ oa18_radiance (rows, columns) float64 159MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ │ oa19_radiance (rows, columns) float64 159MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ │ oa20_radiance (rows, columns) float64 159MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ │ oa21_radiance (rows, columns) float64 159MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ └── DataTree('orphans')
│ Dimensions: (rows: 4092, removed_pixels: 150)
│ Coordinates:
│ altitude (rows, removed_pixels) float32 2MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
│ latitude (rows, removed_pixels) float64 5MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
│ longitude (rows, removed_pixels) float64 5MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
│ Dimensions without coordinates: rows, removed_pixels
│ Data variables: (12/21)
│ oa01_radiance (rows, removed_pixels) float64 5MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
│ oa02_radiance (rows, removed_pixels) float64 5MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
│ oa03_radiance (rows, removed_pixels) float64 5MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
│ oa04_radiance (rows, removed_pixels) float64 5MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
│ oa05_radiance (rows, removed_pixels) float64 5MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
│ oa06_radiance (rows, removed_pixels) float64 5MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
│ ... ...
│ oa16_radiance (rows, removed_pixels) float64 5MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
│ oa17_radiance (rows, removed_pixels) float64 5MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
│ oa18_radiance (rows, removed_pixels) float64 5MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
│ oa19_radiance (rows, removed_pixels) float64 5MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
│ oa20_radiance (rows, removed_pixels) float64 5MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
│ oa21_radiance (rows, removed_pixels) float64 5MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
└── DataTree('quality')
│ Dimensions: (rows: 4092, columns: 4865)
│ Coordinates:
│ latitude (rows, columns) float64 159MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ longitude (rows, columns) float64 159MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ Dimensions without coordinates: rows, columns
│ Data variables: (12/22)
│ oa01_radiance_unc (rows, columns) float64 159MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ oa02_radiance_unc (rows, columns) float64 159MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ oa03_radiance_unc (rows, columns) float64 159MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ oa04_radiance_unc (rows, columns) float64 159MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ oa05_radiance_unc (rows, columns) float64 159MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ oa06_radiance_unc (rows, columns) float64 159MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ ... ...
│ oa17_radiance_unc (rows, columns) float64 159MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ oa18_radiance_unc (rows, columns) float64 159MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ oa19_radiance_unc (rows, columns) float64 159MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ oa20_radiance_unc (rows, columns) float64 159MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ oa21_radiance_unc (rows, columns) float64 159MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
│ quality_flags (rows, columns) uint32 80MB dask.array<chunksize=(1024, 1024), meta=np.ndarray>
└── DataTree('orphans')
Dimensions: (rows: 4092, removed_pixels: 150)
Coordinates:
altitude (rows, removed_pixels) float32 2MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
latitude (rows, removed_pixels) float64 5MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
longitude (rows, removed_pixels) float64 5MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
Dimensions without coordinates: rows, removed_pixels
Data variables: (12/22)
oa01_radiance_unc (rows, removed_pixels) float64 5MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
oa02_radiance_unc (rows, removed_pixels) float64 5MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
oa03_radiance_unc (rows, removed_pixels) float64 5MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
oa04_radiance_unc (rows, removed_pixels) float64 5MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
oa05_radiance_unc (rows, removed_pixels) float64 5MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
oa06_radiance_unc (rows, removed_pixels) float64 5MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
... ...
oa17_radiance_unc (rows, removed_pixels) float64 5MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
oa18_radiance_unc (rows, removed_pixels) float64 5MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
oa19_radiance_unc (rows, removed_pixels) float64 5MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
oa20_radiance_unc (rows, removed_pixels) float64 5MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
oa21_radiance_unc (rows, removed_pixels) float64 5MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
quality_flags (rows, removed_pixels) uint32 2MB dask.array<chunksize=(1024, 150), meta=np.ndarray>
Opening measurement data¶
rad = dt.measurements.oa01_radiance
rad.encoding{'chunks': (1024, 1024),
'preferred_chunks': {'rows': 1024, 'columns': 1024},
'compressor': Blosc(cname='zstd', clevel=3, shuffle=BITSHUFFLE, blocksize=0),
'filters': None,
'_FillValue': 65535,
'scale_factor': 0.013634907081723213,
'add_offset': 0.0,
'dtype': dtype('uint16'),
'coordinates': 'time_stamp latitude longitude'}Underlying data is dask.array
rad.dataLoading...
rad.attrs{'long_name': 'TOA radiance for OLCI acquisition band oa01',
'short_name': 'oa01_radiance',
'standard_name': 'toa_upwelling_spectral_radiance',
'units': 'mW.m-2.sr-1.nm-1',
'valid_max': 65534,
'valid_min': 0}rad.units ## Serving Ambiguity as a replacement for rad.is_scaled
rad.to_masked_array()
rad.sel()Loading...
Simple raster plot¶
Note that using xarray, data is correctly decoded and masked
rad.plot()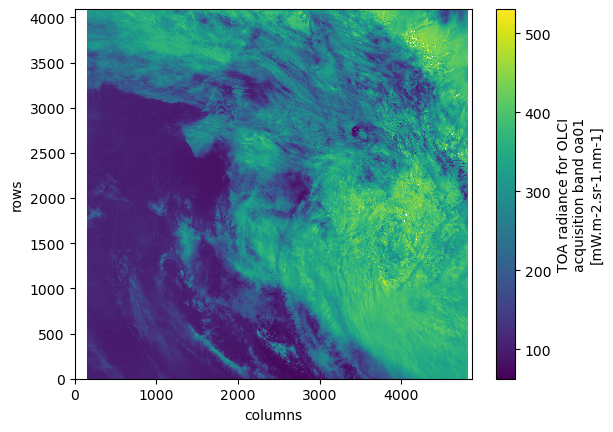
Plot using the coordinates (lon,lat)¶
plt.figure(figsize=(14, 6))
ax = plt.axes()
rad.plot.pcolormesh(ax=ax, x="longitude", y="latitude", add_colorbar=False)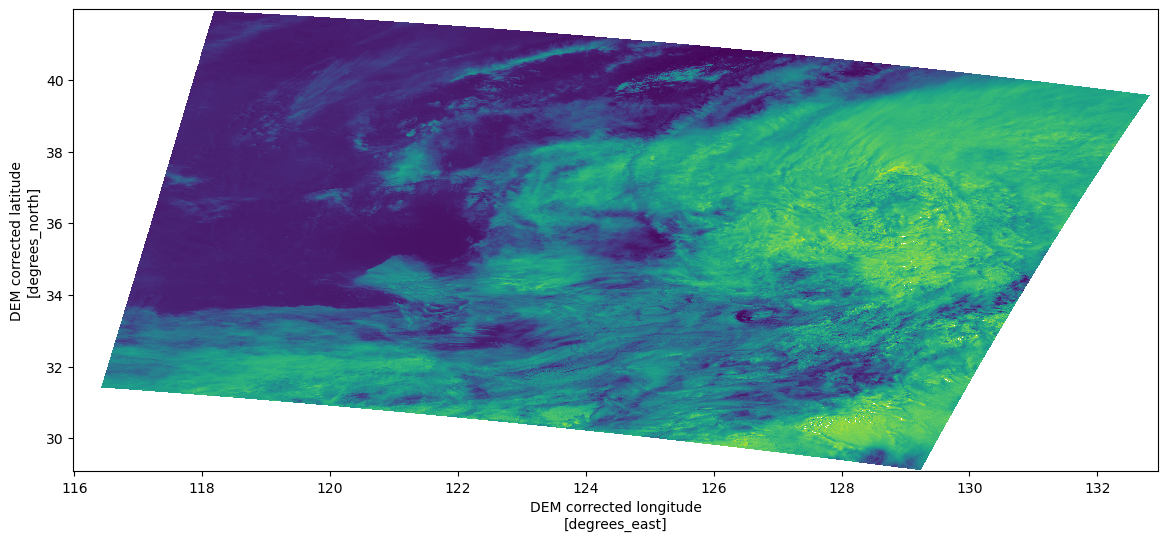
Open Meteorological conditions¶
meteo = dt.conditions.meteorologyInterpolate the atmo. temp. profile at p=832.2 hPa and plot¶
tp = meteo["atmospheric_temperature_profile"].interp(pressure_level=832.2)
tp.plot()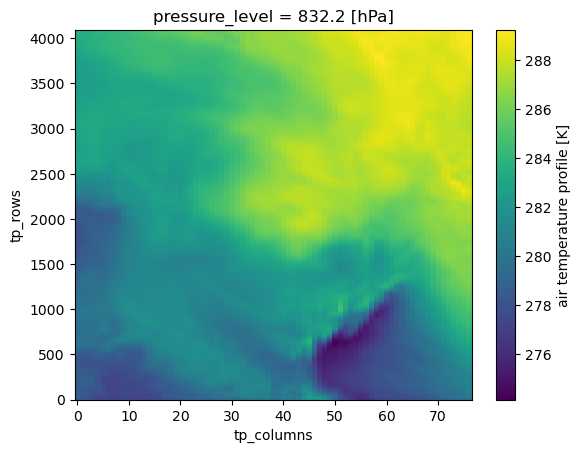
Open sat/sun angles¶
Angles are stored on a tiepoint subgrid
ds = dt.conditions.geometry
dsLoading...
Have a look at condition parameters for removed pixels¶
ds = dt.conditions.orphans
dsLoading...
SLSTR FRP¶
# Define the remote product path
remote_product_path = "https://eopf-public.s3.sbg.perf.cloud.ovh.net/eoproducts/S03SLSFRP_20200908T182648_0179_A298_S883.zarr.zip"
# Construct the fsspec mapper path
store = fsspec.get_mapper(f"zip::{remote_product_path}")
# Load it with datatree
dt = datatree.open_datatree(store, engine="zarr", consolidated=False, chunks={})Opening measurement data (1D)¶
Cannot find the lat/lon in this product ....
meas_in = dt.measurements.inadir
meas_inLoading...
Plot Active Fire pixels positions on a PlateCarree grid¶
import matplotlib.pyplot as plt
fig = plt.figure()
ax = plt.axes(projection=ccrs.PlateCarree())
# ax.set_global()
ax.coastlines()
ax.gridlines(draw_labels=True)
plt.scatter(meas_in.longitude, meas_in.latitude, c=meas_in.frp_mwir, vmax=100)
plt.colorbar()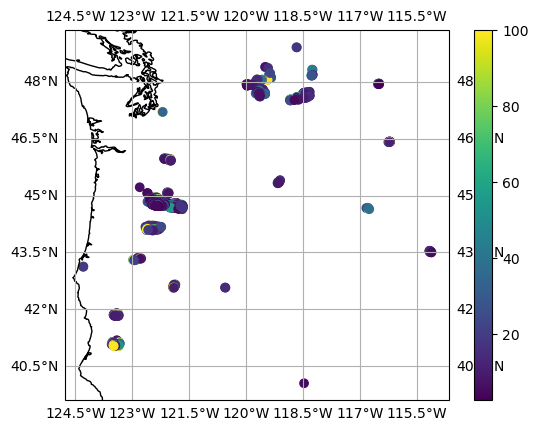
SYN AOD¶
# Define the remote product path
remote_product_path = "https://eopf-public.s3.sbg.perf.cloud.ovh.net/eoproducts/S03SYNAOD_20191227T124211_0060_A109_T883.zarr.zip"
# Construct the fsspec mapper path
store = fsspec.get_mapper(f"zip::{remote_product_path}")
# Load it with datatree
dt = datatree.open_datatree(store, engine="zarr", consolidated=False, chunks={})aod550 = dt.measurements.aod_550
aod550Loading...
aod550.plot()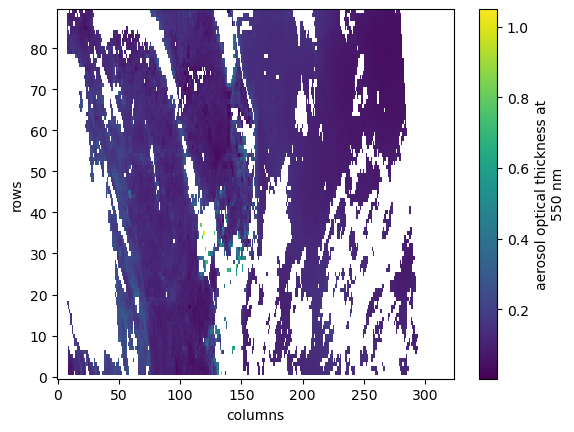
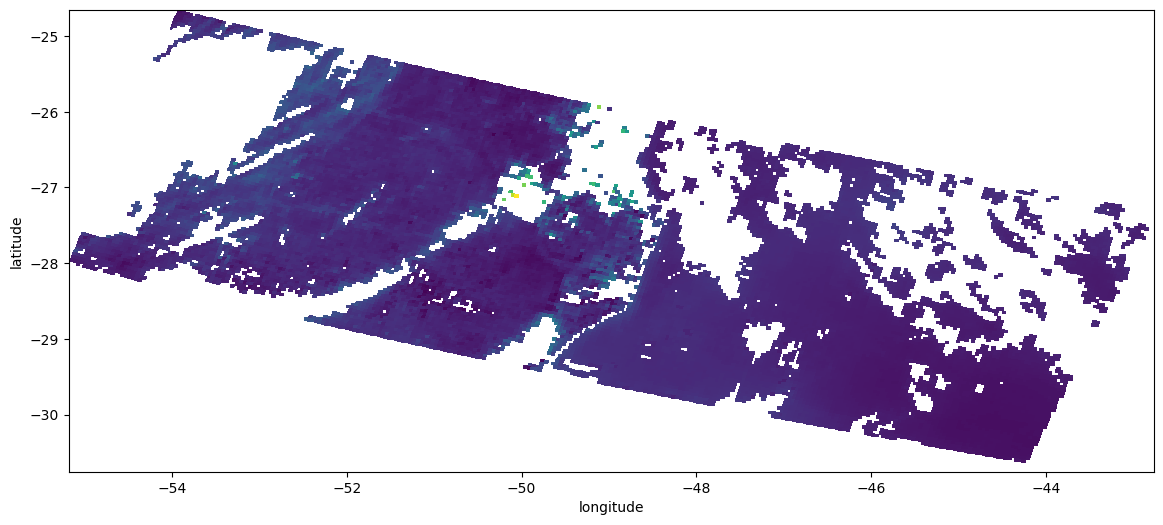
Plot using the coordinates (lon,lat)¶
Note that in SYN AOD product, lat/lon are undefined when the data is missing, which is not correctly handled by matplotlib pcolormesh
# Remove margins
aod550_dropna = aod550.dropna("columns", how="all")
aod550_dropna = aod550_dropna.dropna("rows", how="all")
# Fill remaining missing values
aod550_dropna["latitude"] = (
["rows", "columns"],
aod550_dropna.latitude.bfill("columns").data,
)
aod550_dropna["longitude"] = (
["rows", "columns"],
aod550_dropna.longitude.bfill("columns").data,
)import matplotlib.pyplot as plt
plt.figure(figsize=(14, 6))
ax = plt.axes()
aod550_dropna.plot.pcolormesh(ax=ax, x="longitude", y="latitude", add_colorbar=False)